Abstract
Advances in structural biology have been fueled in part by the development of techniques for large-scale heterologous expression and purification of proteins. However, this step remains a bottleneck in biophysical studies of many proteins. Fusion proteins are often used to increase expression levels, solubility, or both. Here, we compare a recently reported fusion tag, NT*, with maltose-binding protein (MBP), a well-known fusion tag and solubility enhancer. NT* shows high expression and solubility when used as an N-terminal fusion partner for various aggregation-prone peptides.
However, its efficacy in improving the solubility of aggregation-prone globular proteins has not been proven. We found here that although the overall expression levels for the NT* fusions are much higher than those for the MBP fusion, MBP was far superior in improving the solubility of the passenger protein. However, the effective yield after purification of the soluble fraction of the MBP fusion and the NT* fusion was comparable, mainly due to the higher expression levels in the NT* fusion and a smaller weight fraction. net of the passenger protein that is blocked in the fusion. protein.
We conclude that NT* is an excellent fusion tag to enhance the overall expression of globular proteins but does not increase the solubility of the passenger protein compared to MBP. Proteins that are partially soluble or can be refolded in vitro will benefit significantly from N-terminal NT* fusions. However, MBP remains one of the few options for an N-terminal fusion if protein solubility after the expression is critical to preserve its proper folding or activity.
Enhancing the Solubility of Recombinant Proteins Using the MBP Tag
- E. coli expression
Recombinant protein generation requires the use of an expression host and E. coli is often the best choice for this purpose. A combination of simple genetics, ease of cultivation, rapid expression, and high yields make it an attractive host, while a lack of efficient post-translational machinery, bias in codon usage, and difficulty in producing higher molecular weight proteins make it an attractive host. become a disadvantage. However, one of the biggest evils in E.coli expression is an insoluble expression: the phenomenon in which recombinant overexpression results in the formation of insoluble protein aggregates called inclusion bodies.
- Ways to deal with inclusion bodies and improve target protein solubility
When inclusion bodies are formed, scientists often attempt a cumbersome process called in vitro protein solubilization and refolding to recover soluble proteins or attempt to optimize expression in the processor at molecular levels to avoid insolubility issues [recently, there has also been a large amount of interest in the direct use of inclusion bodies as pure and functional protein sources for a variety of biomedical applications.
Examples of process-level optimization include methods that do not require target engineering. For example, optimizing expression conditions, adjusting growth and induction conditions, changing media, buffers, etc. An alternative would be to try a molecular approach and manipulate the target protein by removing undesirable elements within its sequence by creating truncations or mutating certain amino acids. to make the protein more soluble.
You could also take a rational site-directed mutagenesis approach based on structure information or take a directed evolution approach and detect the solubility of random point mutants, deletions and fragments. A better and easier approach would be to adopt a fusion partner strategy and use an upstream fusion partner that would make the target protein more soluble.
- Fusion partner approach to improving protein solubility
Using this approach, a difficult-to-express protein or peptide is fused with another stable, highly soluble protein to stabilize expression, with the idea that the fusion protein will drive the resulting expression. Although many proteins are highly soluble, not all are effective as protein solubility enhancers. In this regard, maltose-binding protein [MBP] may be a very useful protein solubility partner [MBP tag]. In a comparative study aimed at investigating its protein solubility-enhancing properties, the E. coli MBP tag was shown to be a much more effective protein solubility partner than the proteins glutathione-S-transferase (GST) and thioredoxin ( TRx).
About the MBP label and its mechanism of action
MBP is a naturally occurring protein of approximately 42 kDa in E. coli, encoded by the male gene, which is responsible for the absorption, breakdown and transport of maltodextrin, a carbohydrate. Originally developed as a protein expression tag in the 1980s, it is known to significantly improve the solubility of a variety of downstream fused target proteins.
While the exact mechanism of protein solubility enhancement by the N-Terminal MBP tagged recombinant is unclear, it is known to stabilize and protect its downstream passenger protein from proteolytic degradation during and after protein synthesis. Cytoplasmic yields of MBP-fused proteins are also typically higher because the fusion protein provides reliable contexts for efficient translation initiation. It is also proposed that the MBP tag could act as a chaperone through interactions through a solvent-exposed hot spot on its surface that stabilizes the otherwise insoluble passenger protein.
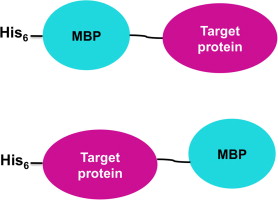
The use of the MBP label
While the MBP tag fused to the N- or C-terminus has been shown to increase recombinant soluble expression, a common approach is to fuse it to the N-terminus. Unfortunately, since no one tag may be ideal for all applications, to increase versatility From labels and downstream applications, combinatorial labelling methods have been developed for various needs.
Two tandemly expressed affinity tags along with a protease cleavage site preceding the target protein allow the use of multiple purification strategies. By incorporating a solubility-enhancing tag in combination with a purification tag, improvements in protein solubility and expression yields are achieved, as well as methods for efficient purification.
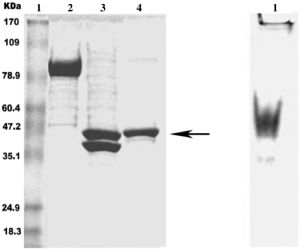
Purification of MBP-tagged proteins
Aside from improved protein solubility, another advantage of using an MBP fusion strategy is to facilitate downstream target protein purification. MBP is a natural affinity tag that binds to amylose resin and can be used for one-step affinity purification by binding to cross-linked amylose. Recombinant MBP-tagged fused proteins bound to immobilized amylose are normally eluted under non-denaturing conditions using maltose [generally 10 mM].
Disadvantages of this strategy
- Cleavage of the target protein can be a problem due to steric hindrance of the MBP tag
- Amylose resin can be brittle and relatively expensive.
- Target protein precipitation after cleavage [often addressed by optimizing protein purification conditions]
- Some MBP fusion proteins do not bind efficiently to amylose resin and even when they do, yields can be an issue [occasionally fixed by optimizing protein purification conditions or changing affinity purification strategy, instead of using instead the His tag]
Conclusion
While there is no simple global solution to address insolubility issues in E.coli expression, a well-designed construct and carefully crafted expression strategy can give you the desired soluble protein. MBP is a reliable solubility label for the production of recombinant soluble proteins, and while solubility is one of the most important elements of a protein production run, solubility should not always be the end goal. Expression levels, protein activity, purity, homogeneity, and stability are important factors that must also be considered before embarking on a recombinant protein production campaign.